There are two main themes in my research. First, most phenotypic evolution is not independent of other phenotypes. Changes in a particular character may influence changes in another and modeling these characters in isolation can mislead our inferences. Second, evolutionary change is heterogeneous. Not all species are going to change the same way at all times and failing to account for that, may again, mislead our inferences. The intersection of these two themes, character dependence and rate heterogeneity, is more natural than it may first appear. This claim is particularly true in the context of phylogenetic comparative methods (PCMs).
‘It is abundantly evident that rates of evolution vary. They vary greatly from group to group, and even among closely related lineages there may be strikingly different rates. Differences in rates of evolution […] are among the reasons for the great diversity of organisms on the earth.’ ‘ - Simpson (1953)
Deep-neural networks as a tool for advancing evolutionary analysis
The anatomy of living species is incredibly diverse and complex. But, can the variety of shapes and forms found in organisms be described by a few basic factors?
By discovering and then studying these potential factors, we may better understand how organisms have changed and evolved over time. To study complex organismal morphology, we use non-linear dimension reduction as a means to discover salient axes of anatomical variation. We utilize various autoencoders and compare their performance to principal component analysis (PCA) for domain specific tasks. Linear dimension reduction performs about as well as non-linear reduction for landmark data. Autoencoders are an exciting way to conduct non-linear dimension reduction because we can actually test hypotheses related to the latent space. There is a lot of potential for improvement: adding phylogeny, better architectures, different data types, explicit structuring of the latent space.
I have recently begun to extend this work into a pure simulation methodology where it can be shown that for non-euclidan shapes, the deep-learning approach exceeds the preformance of PCA and other linear dimension reduction techniques.
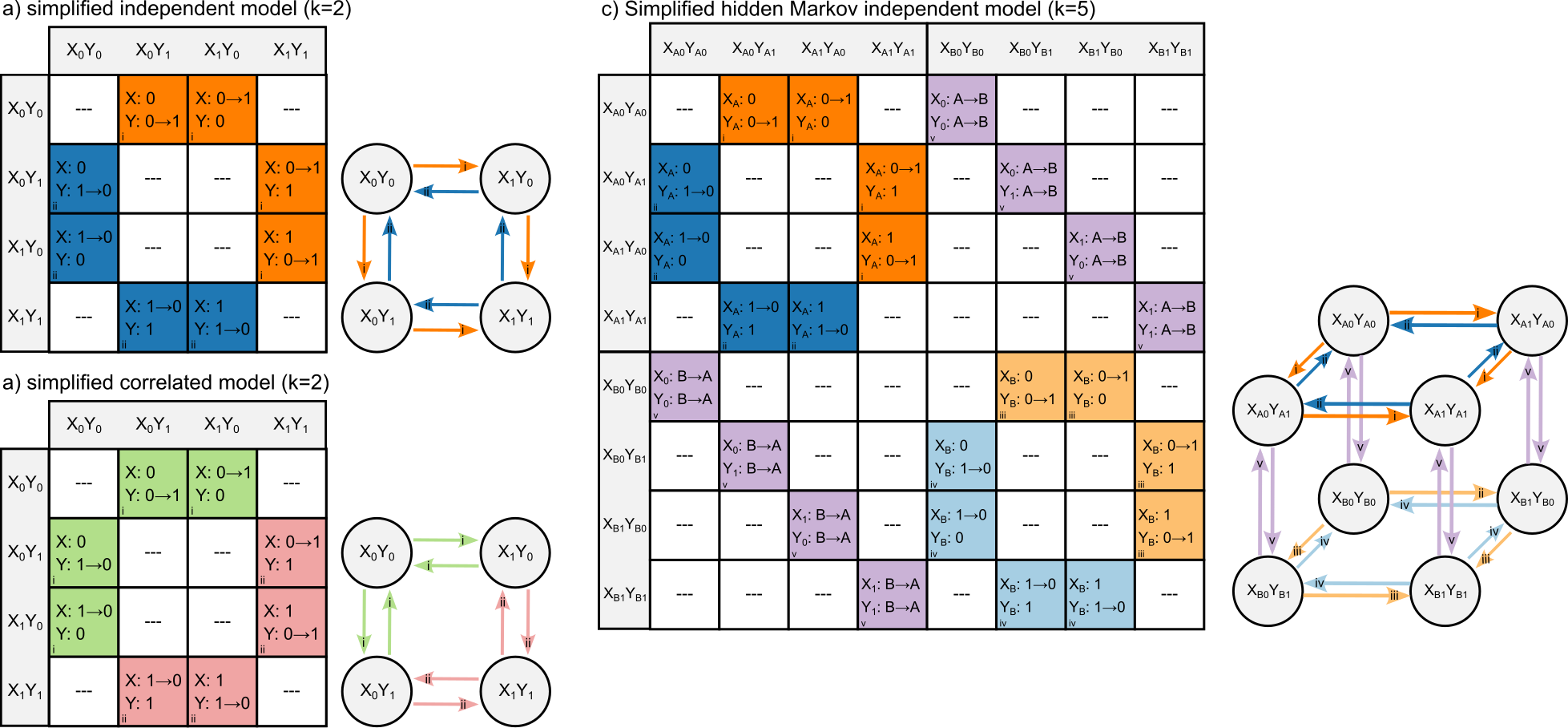
Generalized hidden Markov models
I extended discrete character models to allow for any number of characters with any number of observed or hidden states. This addresses the issue that phenotypic evolution, even when simplified to a discrete character, is better understood as the confluence of several characters evolving together, rather than a single character evolving independently. Furthermore, I demonstrate the some of the advantages of increasing state space from an information theoretic view. By applying this generalization to empirical datasets we drastically alter ancestral state reconstruction finding results more in-line with evo-devo work.
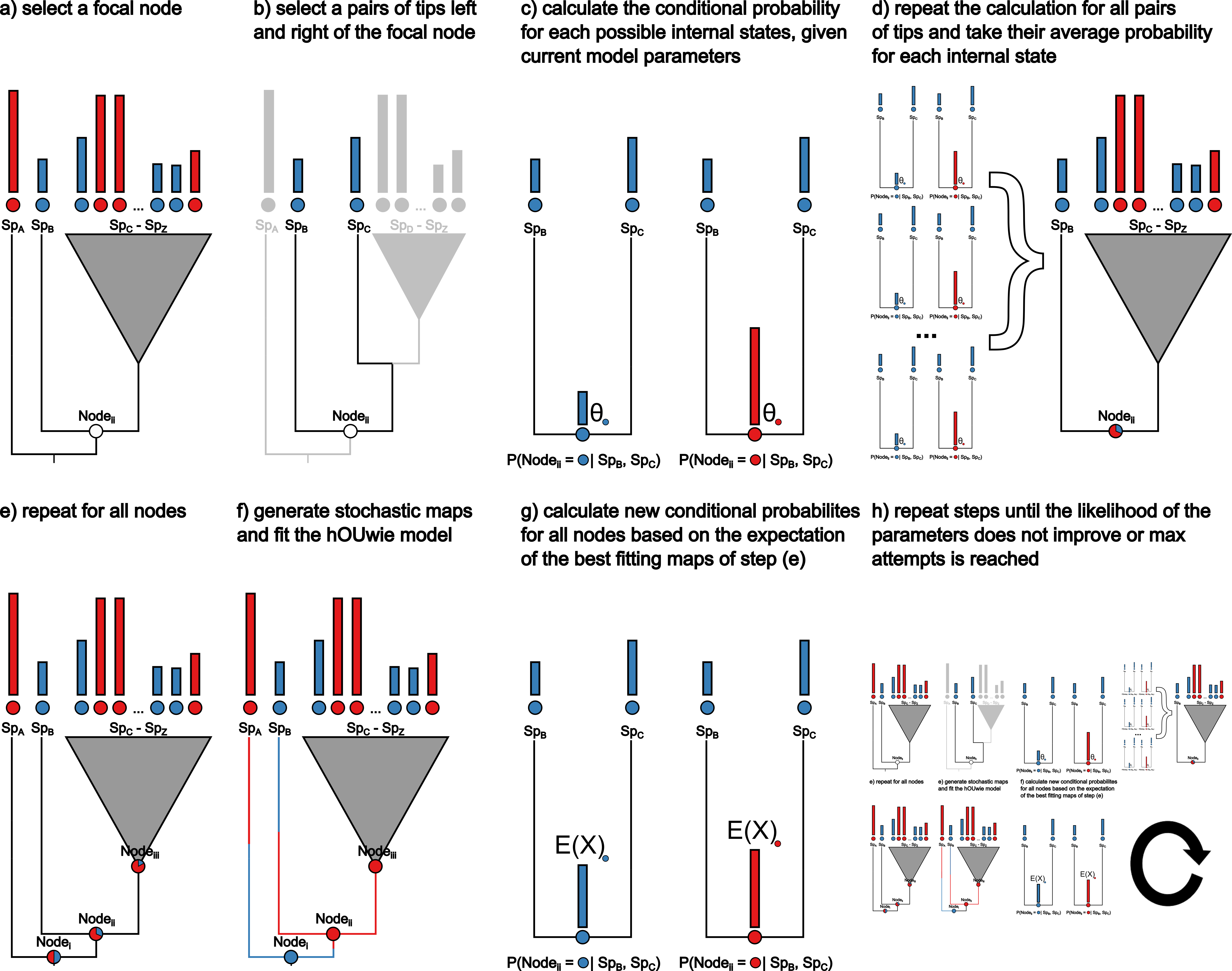
Jointly modeling discrete and continuous characters
I have developed a new model for linking discrete and continuous character evolution. The model called hOUwie, attempts to detect correlation between discrete and continuous characters and estimates their joint evolution. Furthermore, this model is developed with the issues of false correlations in mind and therefore allows for character independent rate heterogeneity as an alternative explanation.
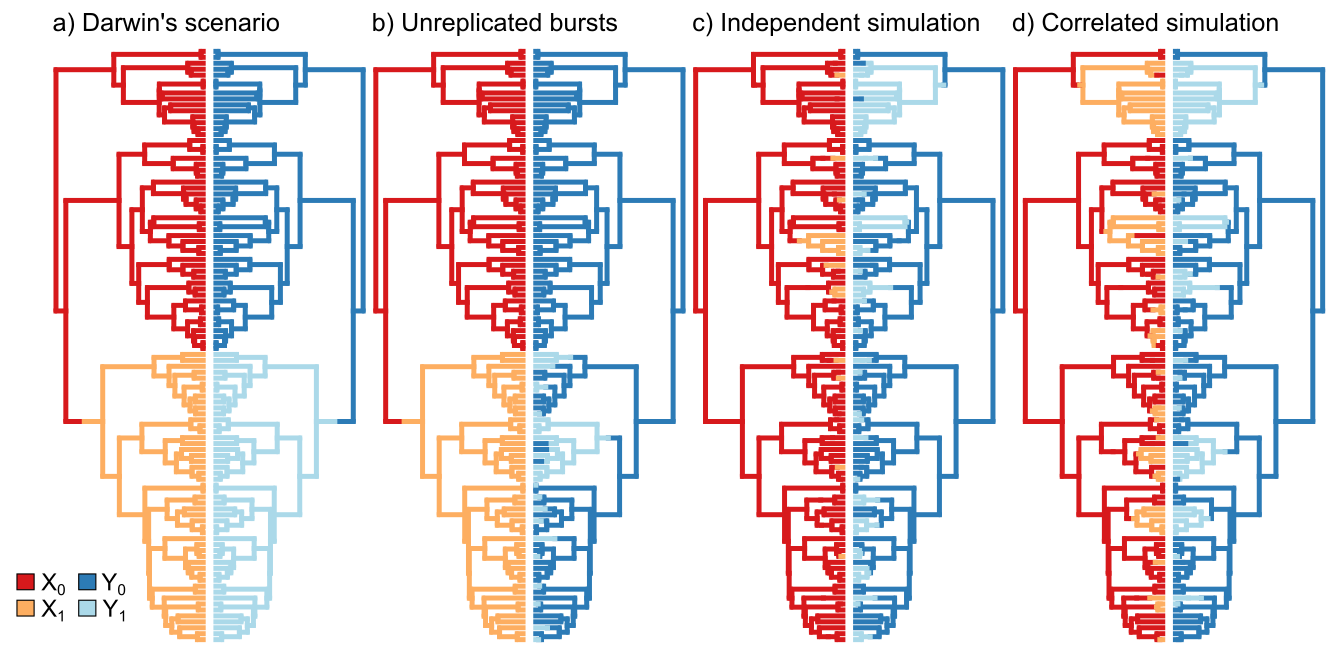
False correlations in PCMs
HMMs are a potential solution to the issue of false correlation between discrete character evolution. Comparative biologists are often interested in whether to discrete characters are correlated with each other as a significant and repeated dependent relationship may give insight into the underlying evolutionary process. However, it has been shown that several commonly used comparative methods are susceptible to false correlations. In this work, I demonstrated that allowing for character independent rate heterogeneity through the application of hidden Markov models, is one way to account for this statistical bias. Nonetheless, there is no amount of methodological massaging that will allow for a satisfying test of macroevolutionary correlation between two synapomorphies.